下载PDF
{"title":"Copy number variant-based genome wide association study reveals immune-related genes associated with parasite resistance in a heritage sheep breed from the United States.","authors":"Zaira M Estrada-Reyes, Ibukun M Ogunade, Andres A Pech-Cervantes, Thomas H Terrill","doi":"10.1111/pim.12943","DOIUrl":null,"url":null,"abstract":"<p><p>Florida Native is a heritage sheep breed in the United States and expresses superior ability to regulate gastrointestinal nematodes. The objective of the present study was to investigate the importance of copy number variants (CNVs) on resistance to natural Haemonchus contortus infections. A total of 300 Florida Native sheep were evaluated. Phenotypic records included fecal egg count (FEC, eggs/gram), FAMACHA© score, percentage cell volume (PCV, %), body condition score (BCS) and average daily gain (ADG, kg). Sheep were genotyped using the GGP Ovine 50K single nucleotide polymorphism (SNP) chip. Log ratios from 45.2 k SNP markers spanning the entire genome were utilized for CNV detection. After quality control, 261 animals with CNVs and phenotypic records were used for the association testing. Association tests were carried out using correlation-trend test and principal component analysis correction to identify CNVs associated with FEC, FAMACHA©, PCV, BCS and ADG. Significant CNVs were detected when their adjusted p-value was <.05 after FDR correction. A total of 8124 CNVs were identified, which gave 246 non-overlapping CNVs. Fourteen CNVs were significantly associated with FEC and PCV. CNVs associated with FEC overlapped 14 Quantitative Trait Locus previously associated with H. contortus resistance. Our study demonstrated for the first time that CNVs could be potentially involved with parasite resistance in Florida Native sheep. Immune-related genes such as CCL1, CCL2, CCL8, CCL11, NOS2, TNF, CSF3 and STAT3 genes could play an important role for controlling H. contortus resistance. These genes could be potentially utilized as candidate markers for selection of parasite resistance in this breed.</p>","PeriodicalId":19931,"journal":{"name":"Parasite Immunology","volume":"44 11","pages":"e12943"},"PeriodicalIF":2.1000,"publicationDate":"2022-11-01","publicationTypes":"Journal Article","fieldsOfStudy":null,"isOpenAccess":false,"openAccessPdf":"https://ftp.ncbi.nlm.nih.gov/pub/pmc/oa_pdf/99/d0/PIM-44-e12943.PMC9786709.pdf","citationCount":"3","resultStr":null,"platform":"Semanticscholar","paperid":null,"PeriodicalName":"Parasite Immunology","FirstCategoryId":"3","ListUrlMain":"https://doi.org/10.1111/pim.12943","RegionNum":4,"RegionCategory":"医学","ArticlePicture":[],"TitleCN":null,"AbstractTextCN":null,"PMCID":null,"EPubDate":"","PubModel":"","JCR":"Q4","JCRName":"IMMUNOLOGY","Score":null,"Total":0}
引用次数: 3
引用
批量引用
Abstract
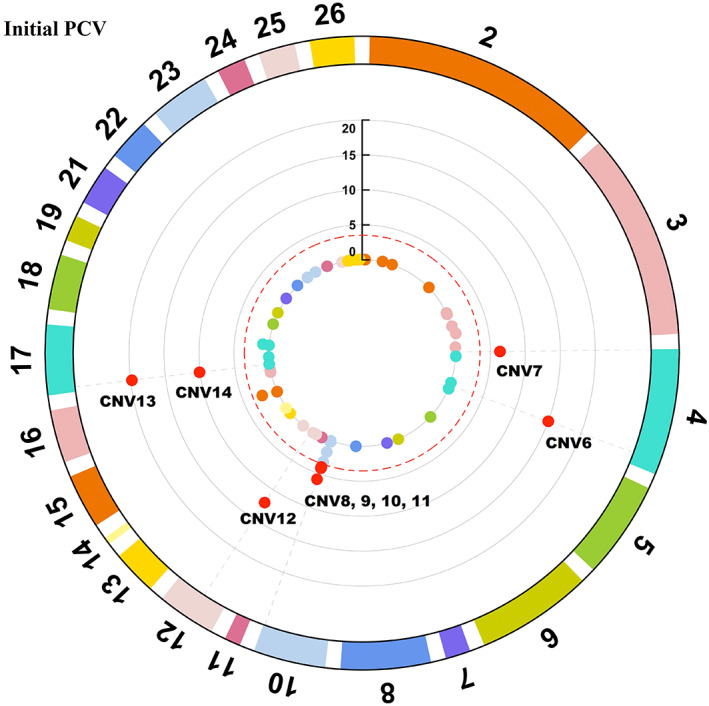
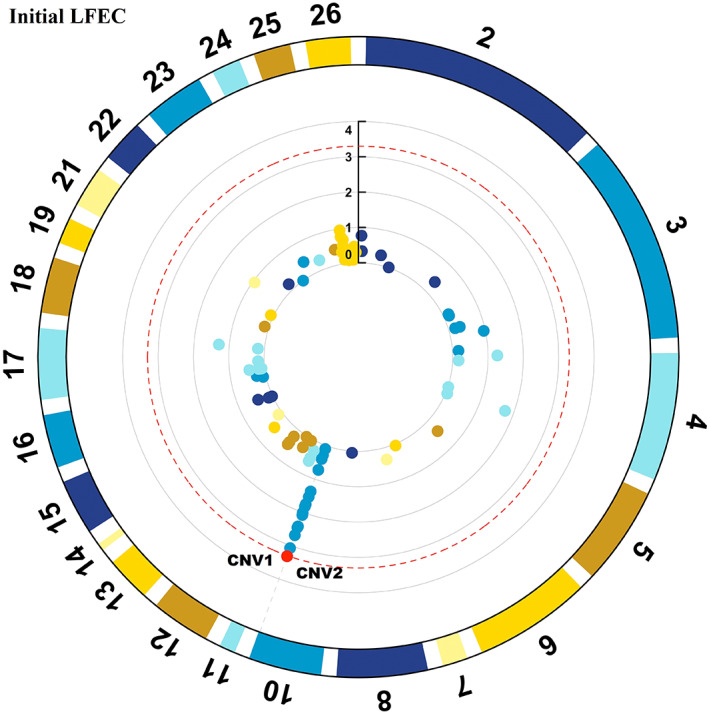
Florida Native is a heritage sheep breed in the United States and expresses superior ability to regulate gastrointestinal nematodes. The objective of the present study was to investigate the importance of copy number variants (CNVs) on resistance to natural Haemonchus contortus infections. A total of 300 Florida Native sheep were evaluated. Phenotypic records included fecal egg count (FEC, eggs/gram), FAMACHA© score, percentage cell volume (PCV, %), body condition score (BCS) and average daily gain (ADG, kg). Sheep were genotyped using the GGP Ovine 50K single nucleotide polymorphism (SNP) chip. Log ratios from 45.2 k SNP markers spanning the entire genome were utilized for CNV detection. After quality control, 261 animals with CNVs and phenotypic records were used for the association testing. Association tests were carried out using correlation-trend test and principal component analysis correction to identify CNVs associated with FEC, FAMACHA©, PCV, BCS and ADG. Significant CNVs were detected when their adjusted p-value was <.05 after FDR correction. A total of 8124 CNVs were identified, which gave 246 non-overlapping CNVs. Fourteen CNVs were significantly associated with FEC and PCV. CNVs associated with FEC overlapped 14 Quantitative Trait Locus previously associated with H. contortus resistance. Our study demonstrated for the first time that CNVs could be potentially involved with parasite resistance in Florida Native sheep. Immune-related genes such as CCL1, CCL2, CCL8, CCL11, NOS2, TNF, CSF3 and STAT3 genes could play an important role for controlling H. contortus resistance. These genes could be potentially utilized as candidate markers for selection of parasite resistance in this breed.
基于拷贝数变异的基因组广泛关联研究揭示了美国一种传统绵羊品种中与寄生虫抗性相关的免疫相关基因。
佛罗里达原生羊是美国的一种传统绵羊品种,具有调节胃肠道线虫的优越能力。本研究的目的是探讨拷贝数变异(CNVs)对天然弯曲血蜱感染抗性的重要性。总共评估了300只佛罗里达本地羊。表型记录包括粪蛋数(FEC,蛋/克)、FAMACHA©评分、细胞体积百分比(PCV, %)、体况评分(BCS)和平均日增重(ADG, kg)。使用GGP羊50K单核苷酸多态性(SNP)芯片对绵羊进行基因分型。利用横跨整个基因组的45.2 k SNP标记的对数比进行CNV检测。经质量控制后,选用261只具有CNVs和表型记录的动物进行关联检测。采用相关趋势检验和主成分分析校正进行关联检验,识别与FEC、FAMACHA©、PCV、BCS和ADG相关的cnv。当调整后的p值为时,检测到显著的CNVs
本文章由计算机程序翻译,如有差异,请以英文原文为准。